Interactions between cells are at the basis of all multicellular life. They allow cells to coordinate decisions such as whether to proliferate, differentiate or die and are therefore essential to generate and maintain healthy tissues. Cell competition regulates survival of cells based on their relative fitness. In heterogenous tissues, weaker cells are removed by surrounding stronger cells. This provides a strong mechanism that governs overall tissue and organismal fitness.
Quality control by cell competition starts during early development where variation in the expression level of the transcription factor Myc causes cell competition-driven cell replacement in the mammalian embryo and heart. In addition, cell competition is an important protector against disease as it aids in the elimination of early malignant cells from tissues and can thereby fulfill a tumorsuppressive function. For example, cells expressing oncogenic Ras are actively eliminated by competitive cell interactions and natural cell competition in the mouse thymus can prevent development of leukemia. But how is cellular fitness determined? And how do cells communicate their fitness to their neighbors? Maybe even more important, how can (external) factors modify cellular fitness? Our group aims to understand how cell competition is regulated by these aspects.
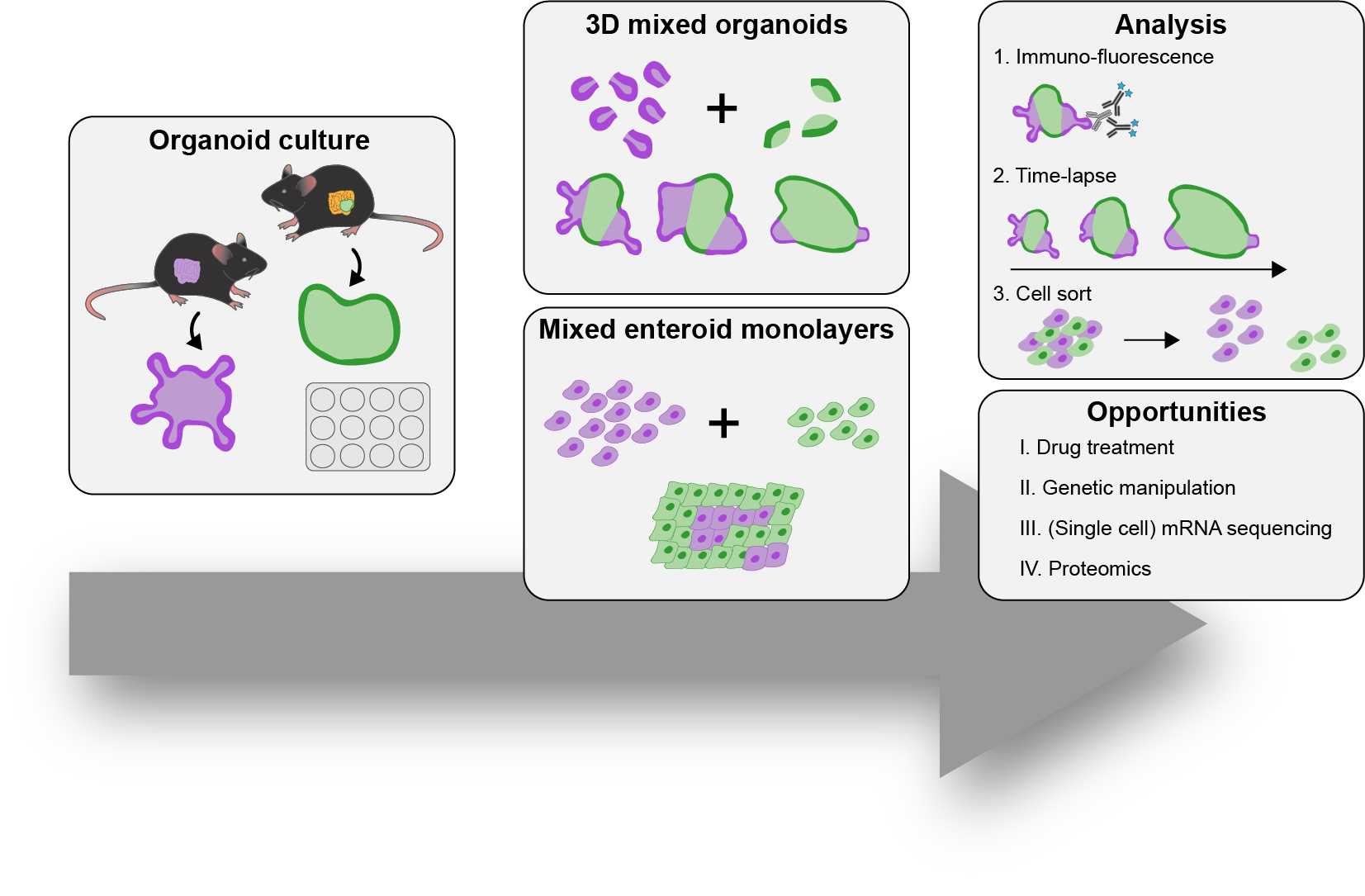
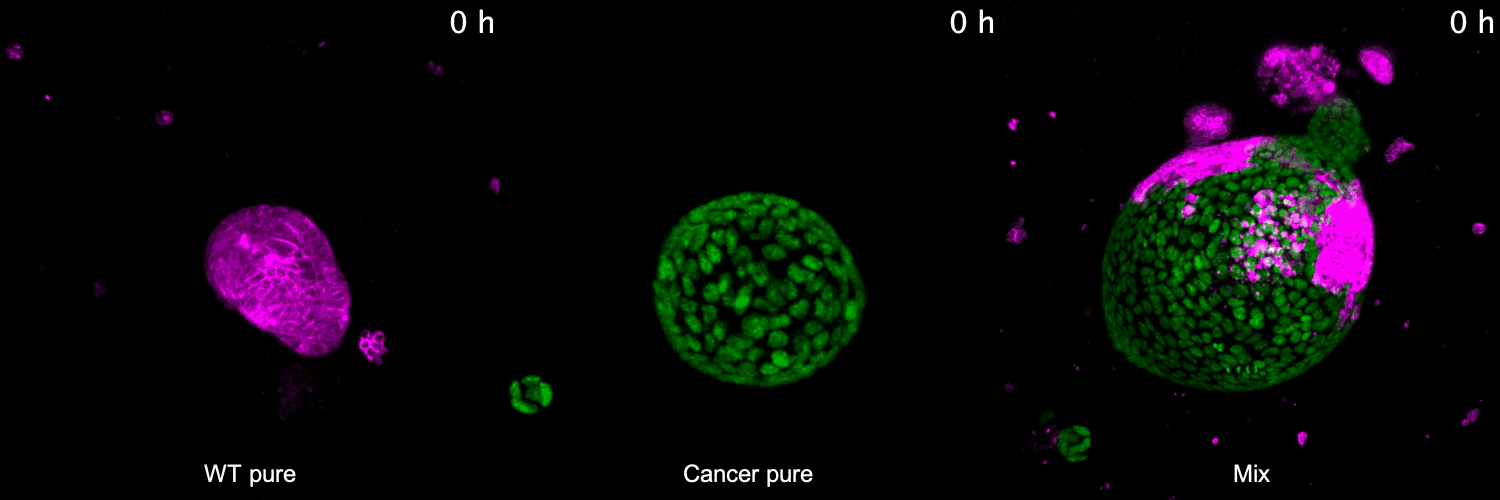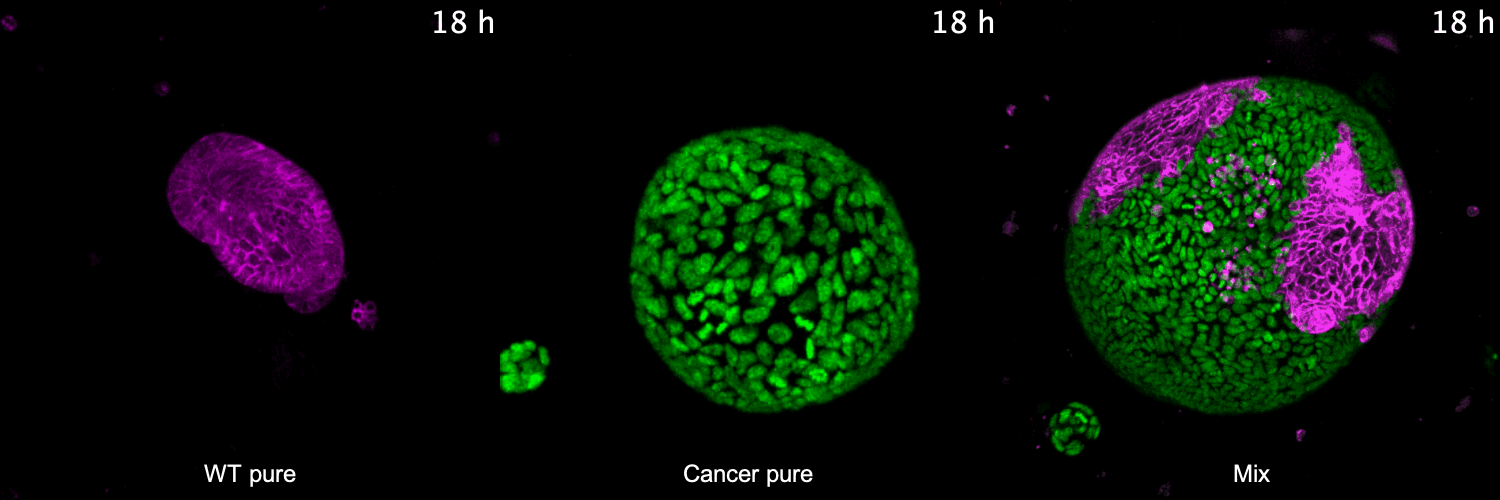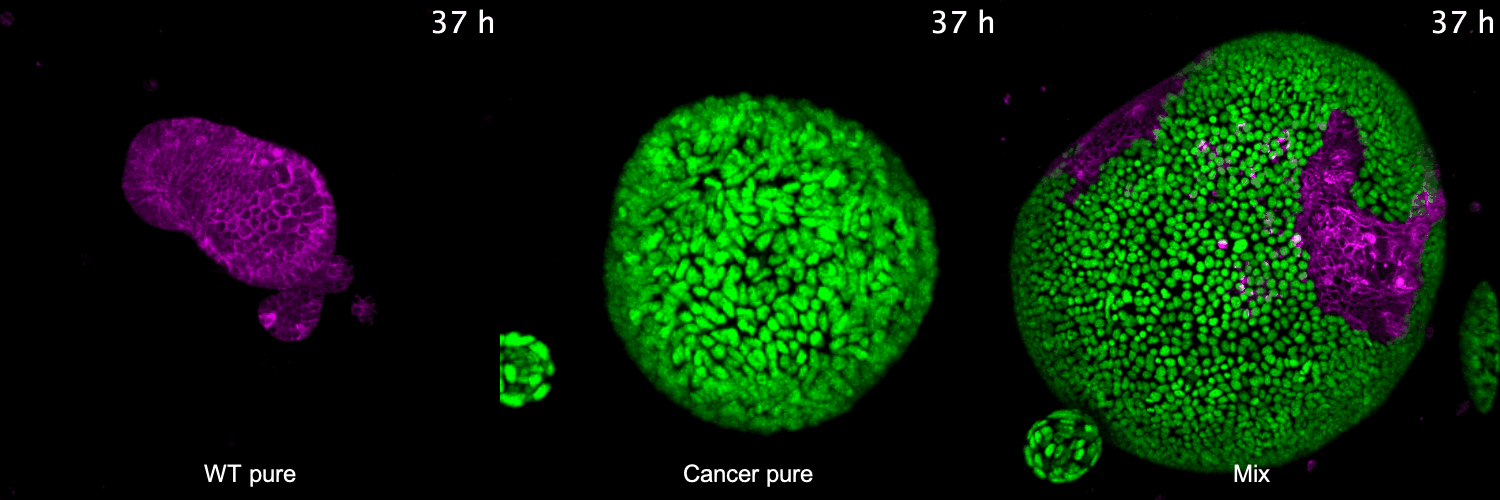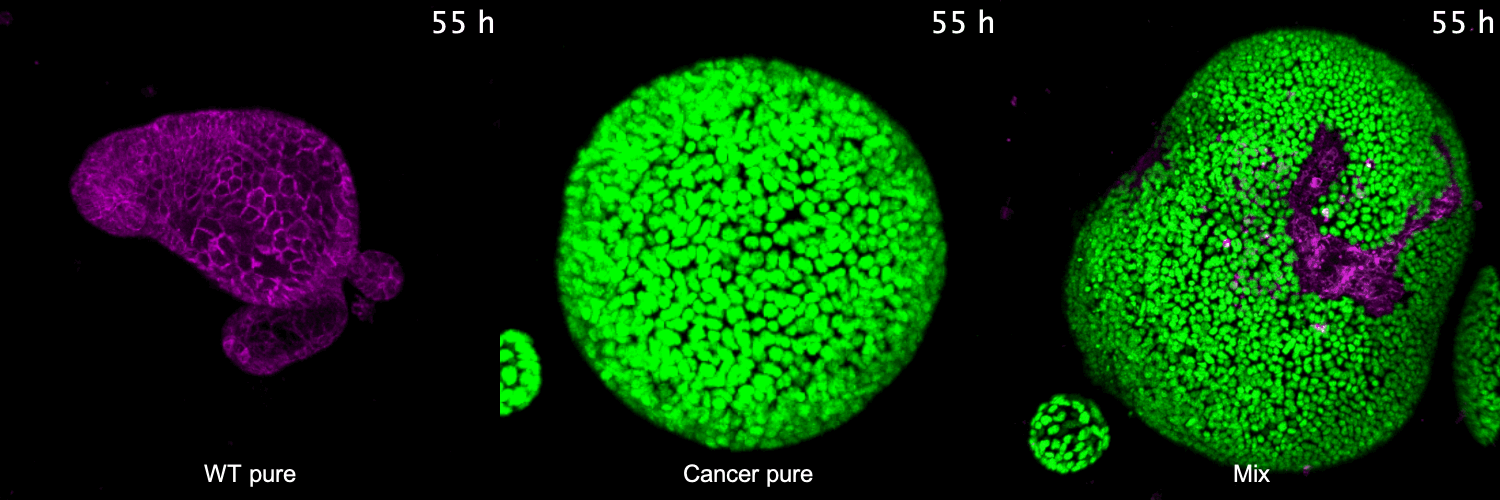
To address these questions, we have developed 3D co-culture systems that allow us to follow the behavior of different cell populations in individual organoids over time. We combine these models with quantitative (time-lapse) microscopy and transcriptome/proteome analysis to unravel the role of cell competition in health disease. We have recently used this system to model the interaction of colorectal cancer with surrounding healthy tissue at the primary and metastatic site. We found that cancer cells use competitive interactions to actively eliminate wild-type intestine cells in enteroid monolayers and 3D organoids. This apoptosis-dependent process boosts proliferation of intestinal cancer cells. The remaining wild-type population activates markers of primitive epithelia and transits to a fetal-like state. JNK signaling is activated in competing cells and is required for cell fate change and elimination of wild-type cells.