Text will be replaced by the actual caption.
We use a combination of systems biology approaches, genetic manipulation, and live-cell imaging to study how epithelial cells polarize, specialize, and become organized into functional tissues.
Cell polarity – the asymmetric distribution of components and functions within a cell along a directional axis – is a fundamental property of cells that is present across the kingdoms of life. Most cells need to distribute proteins, organelles, and functions along an axis of polarity in order carry out their specialized functions and organize the 3D multicellular body.
The most abundant polarized cell type of the animal body is the epithelial cell. Epithelial cells need to establish functionally distinct apical, and basolateral domains to form the epithelial tissues that cover the exterior surfaces of our body and line the surfaces of our organs. Loss of epithelial polarity contributes to epithelial diseases like polycystic kidney disease and retinal dystrophies and contributes to the formation of invasive tumors.
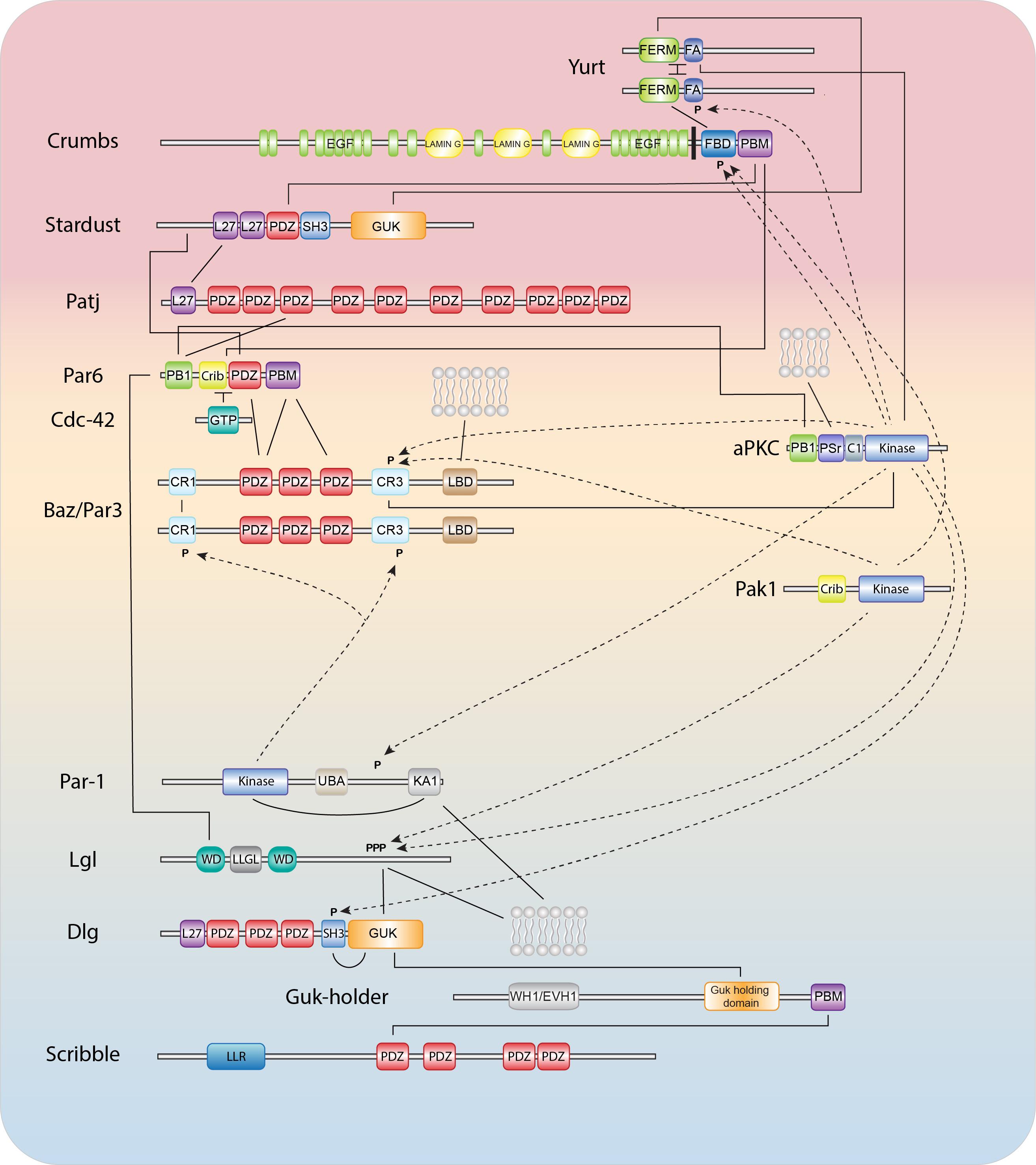
Epithelial polarization and morphogenesis are controlled by an epithelial polarity program in which polarity proteins, protein trafficking, membrane lipid regulators, and actomyosin and microtubule cytoskeletons all contribute important roles (1,2). The establishment of opposing cortical domains is the result of mutually antagonistic interactions and positive feedback loops between a series of evolutionary conserved cortical polarity proteins, most of which were identified through genetic studies in Caenorhabditis elegans and Drosophila melanogaster (3,4). Members of the Partitioning defective (Par) and Crumbs complexes function together in a semi-redundant fashion to establish the apical domain and position cell junctions at the apical/lateral border, while basolateral identity is promoted by the Scribble group proteins, the kinase PAR-1, and the phospho-protein interacting 14-3-3 protein Par5. While the molecular details behind these interactions continue to be worked out, our understanding is still far from complete.
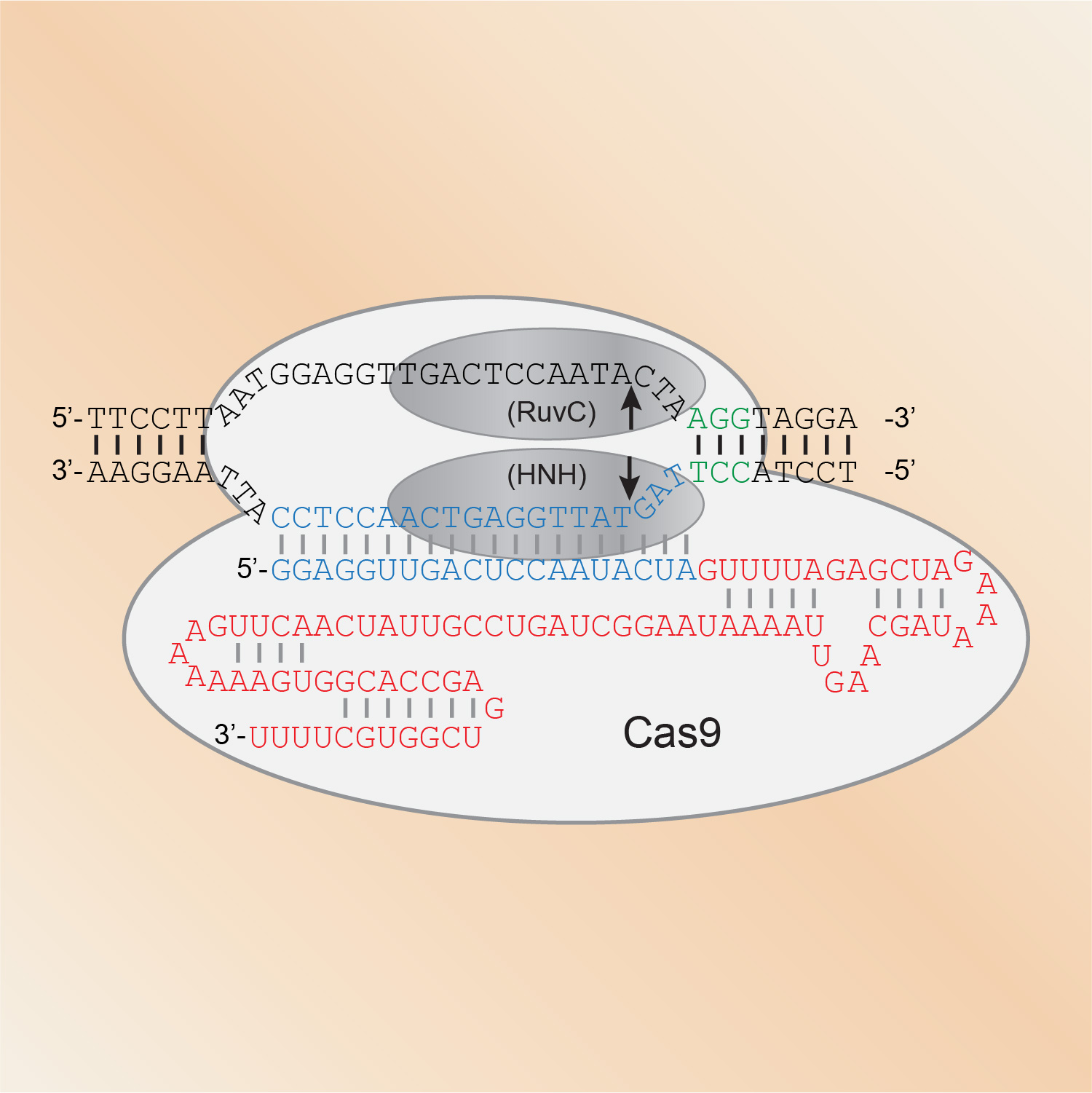
CRISPR/Cas genome engineering has become a staple technology in our lab. Since we showed in 2013 that Cas9 can introduce mutations at specific sites in the genome, we have largely shifted to using CRISPR/Cas9 for our in vivo studies in C. elegans. We use it to express proteins fused to all manner of functional tags, from fluorescent proteins to degron sequences to protein purification tags. We also rely on CRISPR/Cas9 genome engineering to introduce point mutations that for example prevent or mimic phosphorylation, and to generate molecular null alleles by removing entire coding sequences.
Most biological functions are mediated by interactions between proteins, and mapping these interactions is one of the key approaches we use to identify new components involved in polarizing cells and specifying epithelial morphology. We regularly use the yeast two-hybrid system and affinity purification followed by mass spectrometry, both at large and small scale. We also continue to develop or improve protein interaction mapping approaches for C. elegans. For example, we developed a tissue-specific protein purification approach based on biotinylation, and adapted split-intein mediated protein ligation for the detection of interacting protein pairs in C. elegans. Currently, we are working to develop a a fluorescence-based assay to detect protein interaction in vivo in C. elegans.
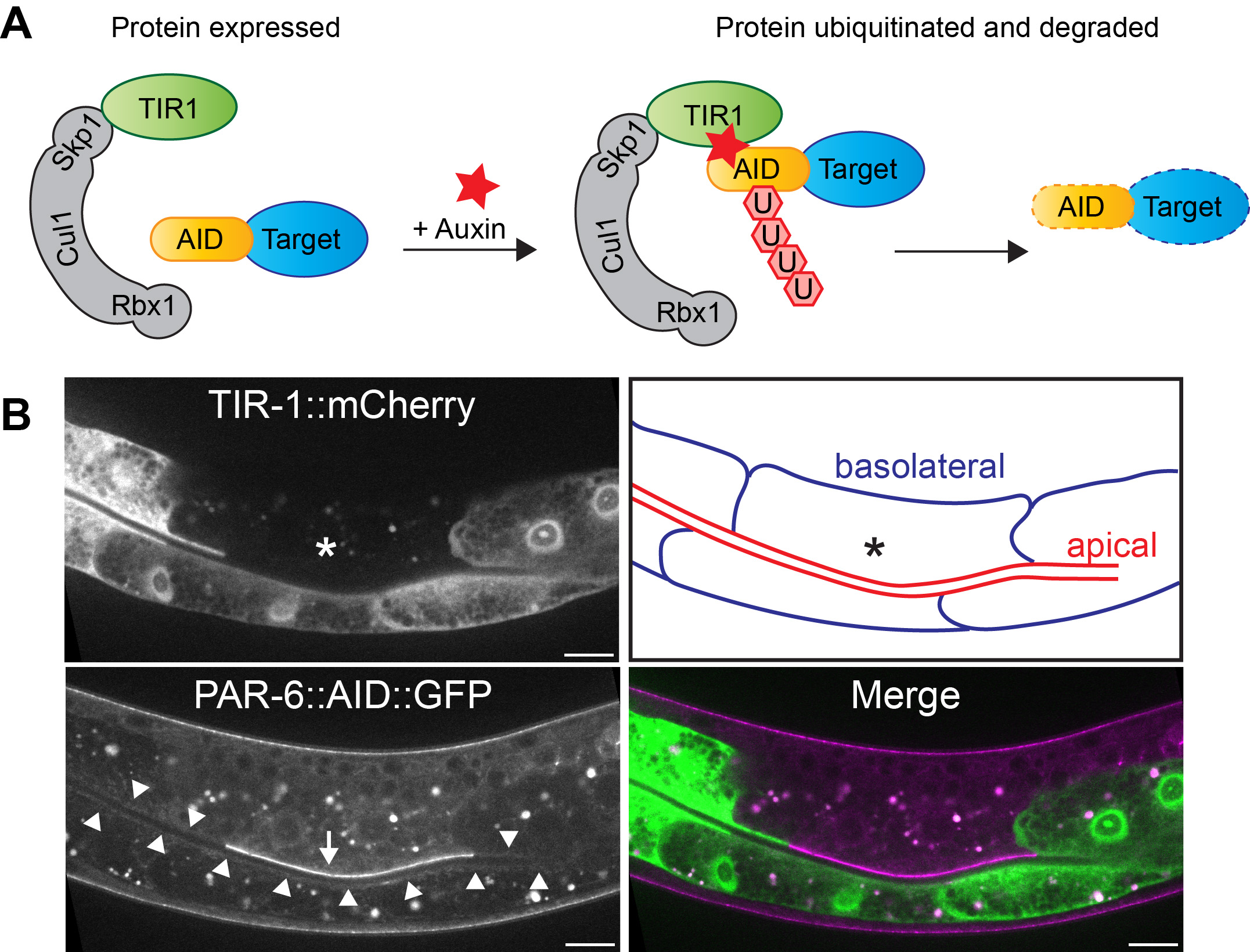
One of our goals is to understand the role of polarity regulators in epithelial tissues throughout development of C. elegans. The problem is, in many cases their loss results in embryonic lethality. To circumvent this, we make use of the auxin-inducible degradation (AID) system. The AID system enables targeted degradation of AID-degron tagged proteins through expression of the plant-derived auxin-dependent E3 ubiquitin ligase specificity factor TIR1 (1,2). By expressing TIR1 from different promoters, both time and tissue specificity are attained. We used this approach for example to uncover novel essential roles for PAR-6/PKC-3 and LET-413 Scribble in the epidermis.
Don’t underestimate the power of a classic mutagenesis screen! Identifying mutants with defects in epithelial polarity or organization remain an attractive approach to identifying genes involved in these processes. And in combination with whole genome sequencing and CRISPR genome engineering, the tedious process of identifying the causative mutation and affected gene is greatly simplified. Through genetics screens, we have identified the intermediate filament regulator BBLN-1 as well as enhancers of crumbs and let-413 mutants.